Enzymatic oligo ligation and cleanup
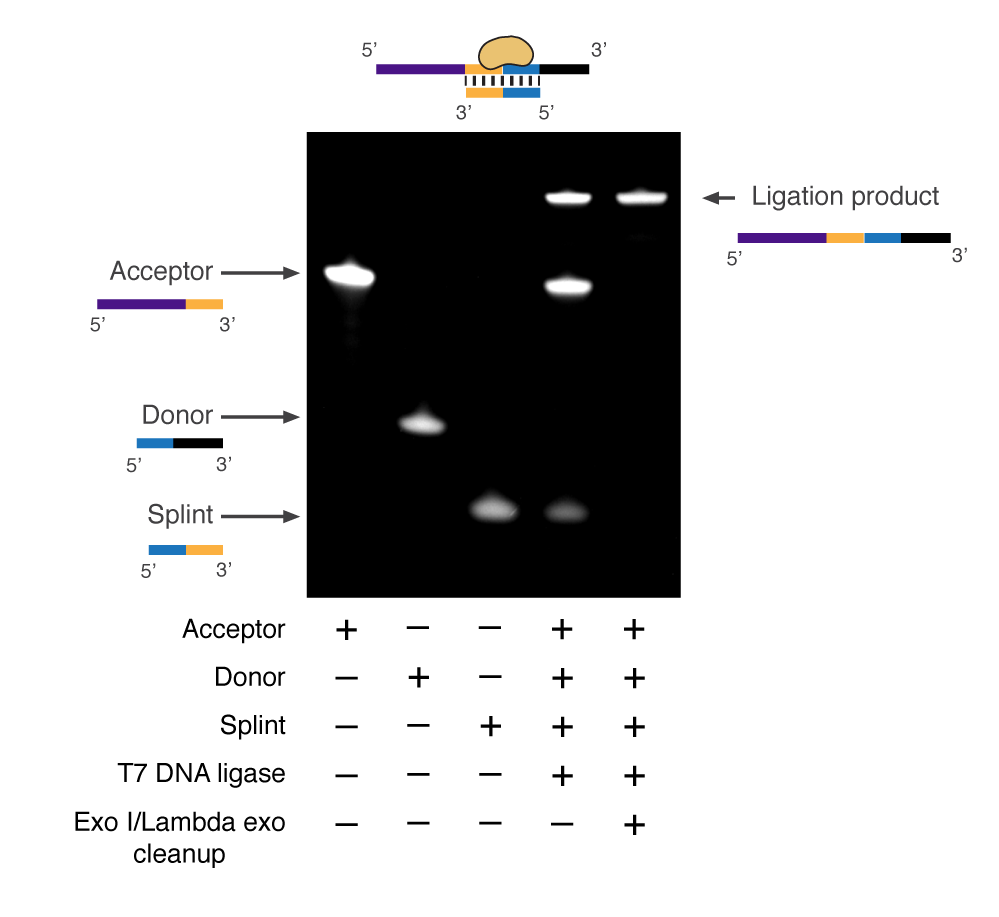
This protocol shows how to use T7 DNA ligase as well as Exonuclease I and Lambda Exonuclease to make longer oligos or combinatorial indexes using a set of building oligos (acceptor, donor and a splint). T7 DNA ligase is used because unlike other ligases it cant support blunt end ligation only sticky end.
The 3'end of the donor oligo is protected against 3'-5' Exo I digestion by phosphorothioate modifications but unreacted donor oligos can still be digested by the 5'phosphate substrate-dependent exonuclease activity by Lambda exonuclease. Unreacted acceptor oligos as well as the splint is digested by 3'-5' activity of Exonuclease I.

🚛 Reagents needed 📦 :
| Item | Article number | Vendor | Distributor |
|---|---|---|---|
| T7 DNA ligase | M0318S | New England Biolabs | BioNordika |
| Exonuclease I (if product <120 nt) | M0293S | New England Biolabs | BioNordika |
| Lambda exonuclease (if product <120 nt) | M0262S | New England Biolabs | BioNordika |
| Oligo Clean & Concentrator (if product <120 nt) | D4060 | Zymo Scientific | Zymo Scientific |
| Monarch PCR cleanup (if product >120nt) | T1030S | New England Biolabs | BioNordika |
| Amicon Ultra-0.5 10k Centrifugal Filters | UFC501024 | Merck | Merck |
Protocol
If the final ligation product is longer than 120 nt then there is no need for enzymatic cleanup the splint and unreacted oligos (size of PCR primers) will be removed by the Monarch PCR cleanup column just like it would remove PCR primers. There is then also no need to include the phosphorothioate modifications on the donor oligo.
If possible use stock concentrations of oligos atleast higher than 400 µM.
1️⃣Mix the oligos and buffer into a hybridization reaction.
| Reagent | Amount | Volume | Final concentration |
|---|---|---|---|
| Nuclease-free water | 4 µl | ||
| Acceptor oligo (5'end of your product) | 0.6 nmol | 1.5 µl | 30 µM |
| Donor oligo (3'end of your product) | 0.4 nmol | 1 µl | 20 µM |
| Splint | 0.6 nmol | 1.5 µl | 30 µM |
| 2X T7 DNA ligase reaction buffer | 10 µl | 1x | |
| TOTAL: | 0.4 nmol | 18 µl (20 µl with enzyme) |
2️⃣ Heat reaction components to 🔥 70°C for 3 min.
3️⃣ Leave reaction components to at room temperature for 5 min for them to anneal 🛬.
4️⃣ Make a diluted stock of T7 DNA ligase by diluting 1:10 and then add 2 µl of the dilution to the reaction to bring it to 20 µl.
5️⃣ Incubate at room temperature for 1h ⏰ to overnight 🌃 🛌.
6️⃣Cleanup reaction with Zymo oligonucleotide spin column:
Add 100 µl Oligo Binding Buffer to 50 µl sample.
Add 400 µl ethanol (95-100%) and mix well by pipetting.
Transfer the 550 µl sample to the Zymo spin column. Spinn at 16,000 rcf for 1 min. Discard the flow through.
Add 750 µl DNA Wash Buffer to the column and centrifuge for 1 min at 16,000 rcf. Discard the flow through.
Dry spinn for 1 min at 16,000 rcf.
Transfer the column to a new tube.
Add 16 µl elution buffer (water or TE) to the center of the column.
Centrifuge for 1 min at 16,000 x g.
Measure the yield on NanoDrop.
If you want to perform enzymatic cleanup do the following otherwise purify with PCR purification kit (Monarch) if product is longer than 150 nt becuase then the column will remove unreacted oligos. Don't load more than 5 µg of oligonucleotides in each exonuclease reaction.
| Reagent | Amount | Volume | Final concentration |
|---|---|---|---|
| T7 DNA ligase reaction | 0.1 nmol | 43 µl | 2 µM |
| 10x Exonuclease I buffer | 5 µl | 1x | |
| Exonuclease I | 20 U | 1 µl | 0.4 U/µl |
| Lambda exonuclease | 5 U | 1 µl | 0.1 U/µl |
| TOTAL: | 50 µl |
7️⃣ Incubate for 2h ⏰ to overnight 🌃 🛌 at 37°C. Heat inactivate the exonucleases by incubating at 80°C for 20 min.
8️⃣ Cleanup with Amicon filter with suitable cut-off (3 kDa for example).
Pool exonuclease treated reactions and fill up with 100 mM NaCl so you have 500 µl to load onto a single Amicon column.
Add the 500 µl sample into the column and spinn at 14'000 rcf for 30 min.
Add 500 µl of 100 mM NaCl onto the column as a wash buffer and spinn for another 30 min at 14'000 rcf .
Revert the Amicon column into a new tube and spinn for 10 min at 1000 rcf.
Measure volume by pipetting and then measure the yield on NanoDrop and then elute further to desired stock concentration (usually 100 µM).